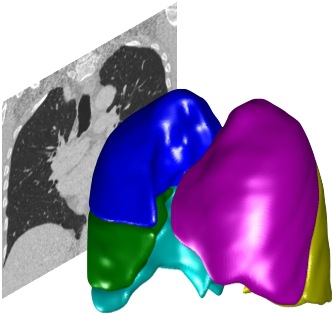
Visualising data¶
First, load a dataset into the PTK GUI.
Once your dataset has loaded, you will a 2D slice from the 3D dataset. What you are seeing in CT is a greyscale image representing the radiodensity of tissue. This is calibrated and (to a good approximation, in general) is proportional to tissue density.
Window/level¶
The first thing to do is set the window and level in order to visualise the features you are interested in. There’s a quick way to do this - select the “Lung” button inside the Window/Level presets pane at the right of the screen. Try out other presets.
Now try using the window/level tool. Note the W/L button underneath the image is selected (if it isn’t, select it now). Click inside the lung image and drag the mouse around. Note how the values of the window and level sliders below the image change. Select different Window/Level presets from the presets panel and see how the numbers change again. You can manually edit the window and level values if you want specific values.
Moving through image slices¶
There are several ways to move through the images slices in the 3D image:
Scroll the mousewheel while over the image
up/down cursors on keyboard
scrollbar to the right of the image
Select the Cine tool (
nkey) (in the toolbar under the image), and drag up/down inside the image
Image orientation¶
The three available image orientations are Coronal, Sagittal and Axial. Use the buttons at the bottom-left of the screen, or the keys C, S, A to change orientation
Density values¶
As you move the mouse cursor over the image, the value of the image under the cursor is displayed underneath the image (after the coordinate values). For CT, two values are shown; the raw image value (I:xxx) and the calibrated Hounsfield value (HU:xxx).